Publications
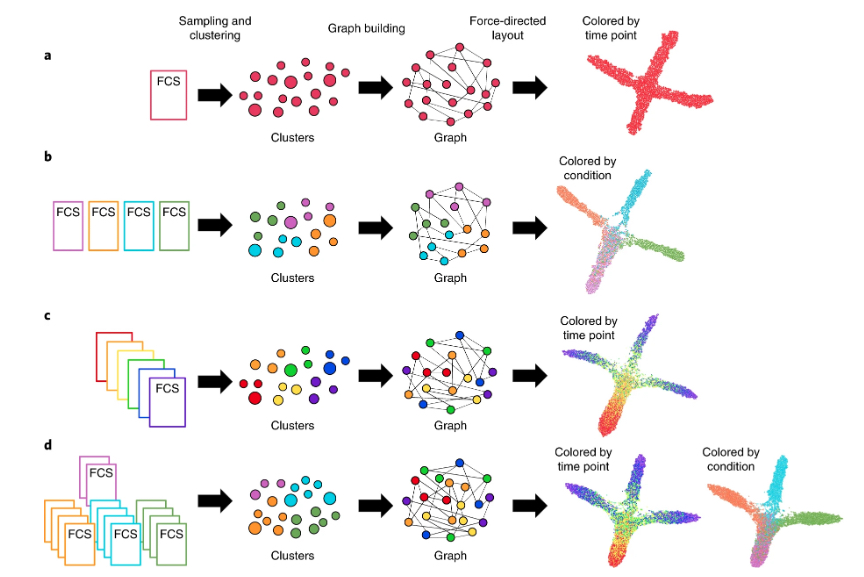
Melissa E. Ko*, Corey M. Williams*, Kristen I. Fread, Sarah M. Goggin, Rohit S. Rustagi, Gabriela K. Fragiadakis, Garry P. Nolan & Eli R. Zunder. FLOW-MAP: a graph-based, force-directed layout algorithm for trajectory mapping in single-cell time course datasets. Nature Protocols. 2020. *contributed equally
- PMID:
- Full Text Additional Links
- FLOW-MAP: a graph-based, force-directed layout algorithm for trajectory mapping in single-cell time course datasets
- FLOW-MAP code and GUI
Access the paper
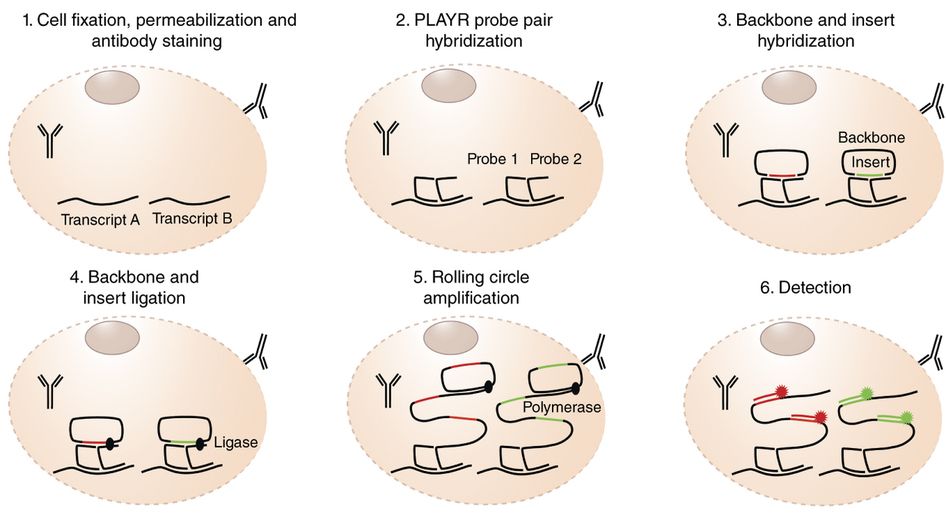
Frei AP*, Bava FA*, Zunder ER, Hsieh EW, Chen SY, Nolan GP, Gherardini PF. Highly multiplexed simultaneous detection of RNAs and proteins in single cells. Nature Methods. 2016. *contributed equally
- PMID: 26808670
- Full Text Additional Links
- Nolan Lab
- PLAYRDesign Software
- Nature Education Bio 2.0 Blog: Rapid RNA analysis at a single cell level: a PLAYR's story
Access the paper
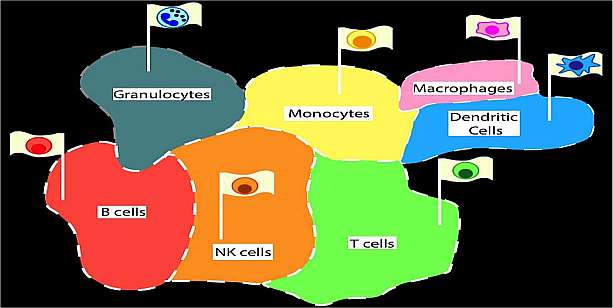
Spitzer MH*, Gherardini PF*, Fragiadakis GK, Bhattacharya N, Yuan RT, Hotson AN, Finck R, Carmi Y, Zunder ER, Fantl WJ, Bendall SC, Engleman EG, Nolan GP. A dynamic immune system reference map: system-wide organization with functional correlates. Science. 2015. *contributed equally
- PMID: 26160952
- Full Text
- Cytobank Report, including downloadable FCS files Additional Links
- Engleman Lab
- Nolan Lab
- Sciencescape: Mass Cytometry: A First Time Glimpse at the Dynamic Immune System
Access the paper
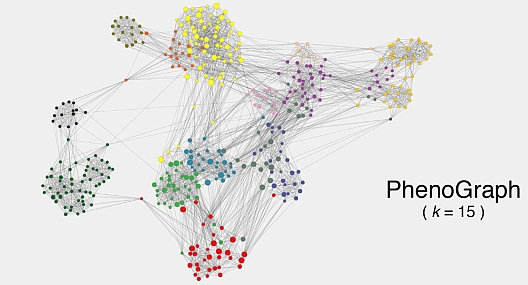
Levine JH*, Simonds EF*, Bendall SC*, Davis KL, Amir ED, Litvin O, Fienberg H, Jager A, Pinkus L, Zunder ER, Finck R, Gedman AL, Radtke I, Downing JR, Pe’er D, Nolan GP. Data-Driven Phenotypic Dissection of AML Reveals Progenitor-like Cells that Correlate with Prognosis. Cell. 2015. *contributed equally

- PMID: 26095251
- Full Text
- Cytobank Report, including downloadable FCS files Additional Links
- Pe'er Lab
- Nolan Lab
- From mass cytometry to cancer prognosis
Access the paper
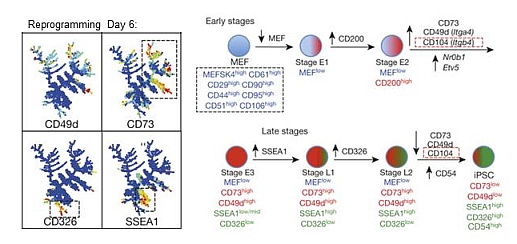
Lujan E*, Zunder ER*, Nolan GP, Wernig M. Early regulators of iPSC reprogramming identified by a novel set of transient intermediates. Nature. 2015. *contributed equally
- PMID: 25830878
- PMCID: PMC4441548
- Full Text Additional Links
- Wernig Lab
- Nolan Lab
- Stem Cell Portal: Reprogramming Revealed Using a Mass-terly Strategy
Access the paper
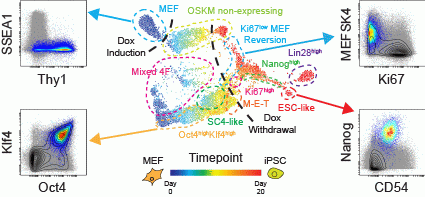
Zunder ER*, Lujan E*, Goltsev Y, Wernig M, and Nolan GP. A Continuous Molecular Roadmap to iPSC Reprogramming Through Progression Analysis of Single Cell Mass Cytometry. Cell Stem Cell. 2015. *contributed equally
- PMID: 25748935
- PMCID: PMC4401090
- Full Text
- Deposited GEO Dataset: GSE56764
- Cytobank Report, including downloadable FCS files Additional Links
- Nolan Lab
- Wernig Lab
- Cell Stem Cell: Charting a map through the cellular reprogramming landscape
Access the paper
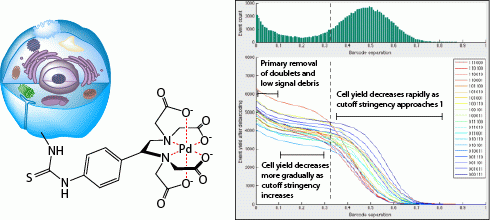
Zunder ER*, Finck R*, Behbehani GK, Amir ED, Krishnaswamy S, Gonzalez VD, Lorang CG, Bjornson Z, Spitzer MH, Bodenmiller B, Fantl WJ, Pe’er D, Nolan GP. Palladium-based Mass-Tag Cell Barcoding with a Doublet-Filtering Scheme and Single Cell Deconvolution Algorithm. Nature Protocols. 2015. *contributed equally
- PMID: 25612231
- PMCID: PMC4347881
- Full Text
- Cytobank Report, including downloadable FCS files Additional Links
- Nolan Lab
- Pe'er Lab
- Single-Cell Debarcoding Software
- Single-Cell Debarcoding Manual
- Fluidigm Spotlight: Exploring Impossible Science
- Fluidigm Cell-ID 20-Plex Pd Barcoding Kit
Access the paper
Behbehani GK, Thom C, Zunder ER, Finck R, Gaudilliere B, Fragiadakis GK, Fantl WJ, Nolan GP. Transient partial permeabilization with saponin enables cellular barcoding prior to surface marker staining. Cytometry Part A. 2014.

- PMID: 25274027
- PMCID: PMC4361015
- Full Text
- Cytobank Report, including downloadable FCS files Additional Links
- Nolan Lab
Access the paper
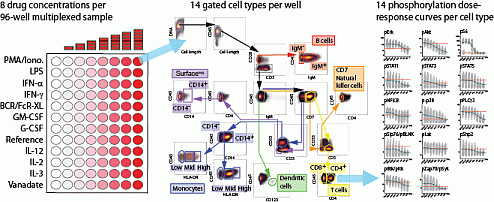
Bodenmiller B*, Zunder ER*, Finck R*, Chen TJ, Savig ES, Bruggner RV, Simonds EF, Bendall SC, Sachs K, Krutzik PO, Nolan GP. Multiplexed mass cytometry profiling of cellular states perturbed by small-molecule regulators. Nature Biotechnology. 2012. *contributed equally
- PMID: 22902532
- PMCID: PMC3627543
- Full Text
- Cytobank Report, including downloadable FCS files Additional Links
- Nolan Lab
- Nature Methods: Research Highlights
- The Scientist: No-Mo-Slow-Flow
Access the paper
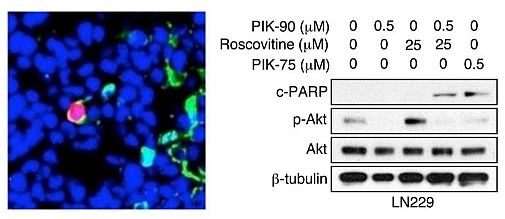
Cheng CK, Gustafson WC, Charron E, Houseman BT, Zunder ER, Goga A, Gray NS, Pollok B, Oakes SA, James CD, Shokat KM, Weiss WA, Fan QW. Dual blockade of lipid and cyclin-dependent kinases induces synthetic lethality in malignant glioma. Proceedings of the National Academy of Sciences, USA. 2012.
- PMID: 22802621
- PMCID: PMC3411950
- Full Text Additional Links
- Weiss Lab
- Shokat Lab
Access the paper
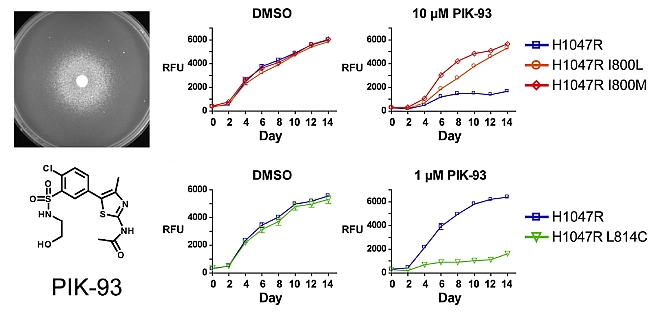
Zunder ER, Knight ZA, Houseman BT, Apsel B, Shokat KM. Discovery of drug-resistant and drug-sensitizing mutations in the oncogenic PI3K isoform p110alpha. Cancer Cell. 2008.
- PMID: 18691552
- PMCID: PMC2720137
- Full Text Additional Links
- Shokat Lab
- Cancer Cell: Drug-resistant phosphatidylinositol 3-kinase: guidance for the preemptive strike
- HHMI: Studies Aim to Preempt Resistance to New Class of Cancer Drugs
Access the paper
Knight ZA, Gonzalez B, Feldman ME, Zunder ER, Goldenberg DD, Williams O, Loewith R, Stokoe D, Balla A, Toth B, Balla T, Weiss WA, Williams RL, Shokat KM. A pharmacological map of the PI3-K family defines a role for p110alpha in insulin signaling. Cell. 2006.

- PMID: 16647110
- PMCID: PMC2946820
- Full Text
- Deposited Structures: 2CHW, 2CHX, 2CHZ Additional Links
- Shokat Lab
- Williams Lab
- Weiss Lab
- Balla Lab
- Stokoe Lab
- Loewith Lab
- Cell: Using isoform-specific inhibitors to target lipid kinases
- HHMI: Chemical Biology Suggests New Way to Thwart Brain Cancer
- Faculty of 1000 Evaluation
Access the paper